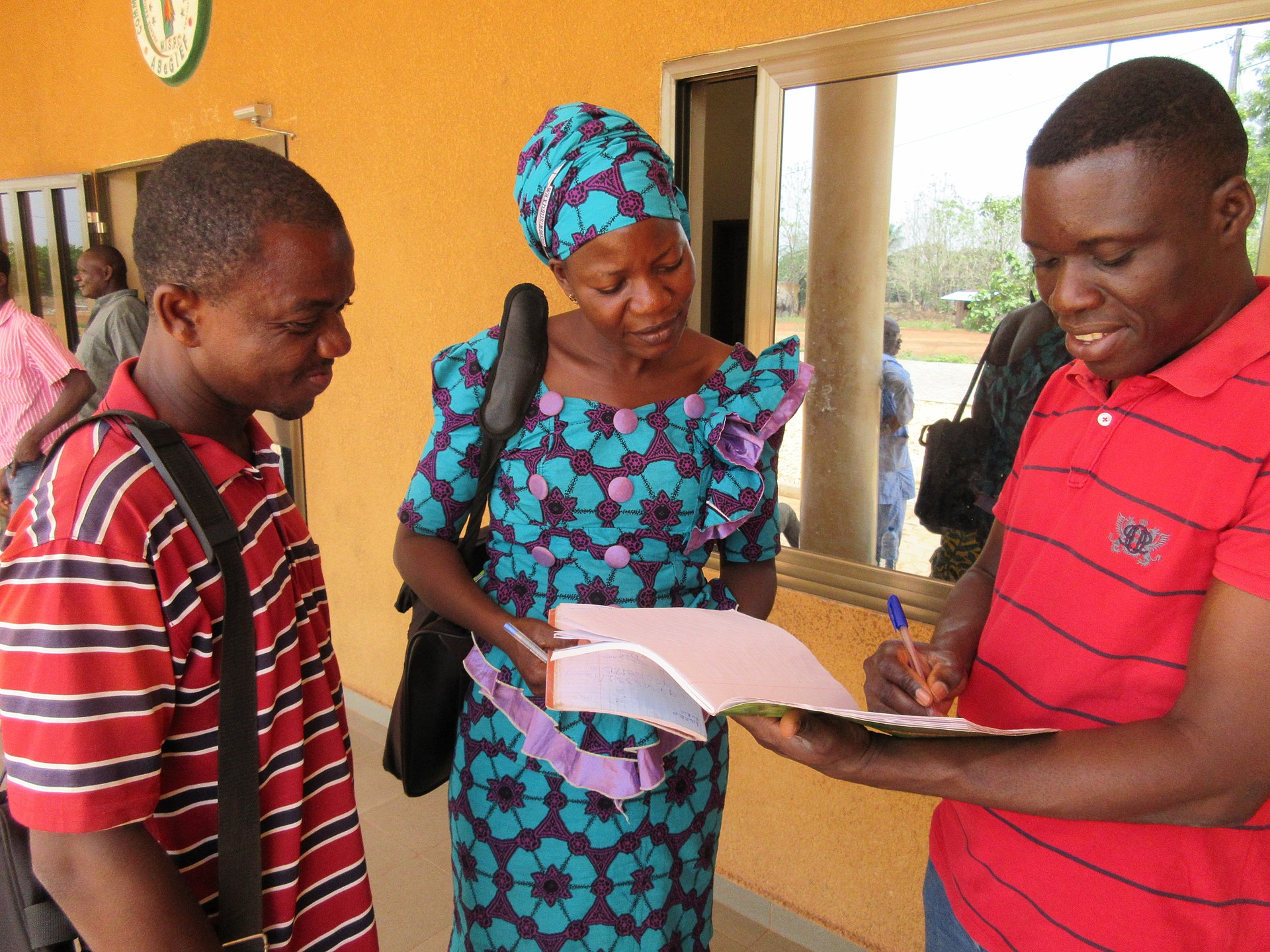

Lassa fever is a viral haemorrhagic disease endemic to West Africa, caused by the Lassa virus and primarily transmitted through exposure to food or surfaces contaminated by infected rodents. Human-to-human transmission can also occur, particularly in healthcare settings without adequate infection control. Symptoms range from mild fever and weakness to severe complications such as bleeding, organ failure, and, in some cases, death.
Nigeria continues to experience seasonal outbreaks, and recent data from the Nigeria Centre for Disease Control and Prevention shows that 2026 is following a concerning trajectory.
As of Epidemiological Week 8 (February 2026):
- 404 confirmed cases have been recorded
- 99 deaths, with a case fatality rate (CFR) of 24.5%
- Cases have spread across 18 states and 67 Local Government Areas
While weekly confirmed cases have slightly declined (77 new cases in Week 8 compared to 82 in Week 7), the rising fatality rate higher than 18.8% in 2025 signals ongoing gaps in early detection and treatment access.
More recent updates suggest the trend persists. By March 2026, Nigeria had recorded over 580 confirmed cases and more than 140 deaths, maintaining a high fatality rate above 24%.
Geographic and Demographic Trends
The outbreak remains highly concentrated:
- Bauchi (30%), Ondo (21%), and Taraba (19%) account for the majority of cases
- Together with Edo and Benue, these states contribute 84% of all confirmed infections
The most affected group is young adults aged 21–30, although cases span from infants to older adults. Men are slightly more affected than women (ratio 1:0.8).
Healthcare workers also remain at risk, with new infections recorded among frontline responders highlighting persistent vulnerabilities in infection prevention systems.
Treatment Gaps and a New Breakthrough
Currently, treatment options for Lassa fever are limited. The antiviral drug ribavirin is widely used but is most effective only when administered early, which is often not the case due to delayed diagnosis.
However, new research published in Nature introduces a promising development. Scientists have identified an oral antiviral drug, 4′-fluorouridine (4′-FIU), that demonstrated strong efficacy in treating Lassa fever in nonhuman primates.
Key findings from the study include:
- Effectiveness even when treatment begins several days after infection
- Significant reduction in viral load
- Improved survival outcomes
This is particularly important because Lassa fever is often diagnosed late, when existing treatments are less effective. An oral drug also offers practical advantages for deployment in rural and resource-limited settings.
What This Means for Public Health in Africa
The combination of rising fatality rates and continued geographic spread underscores the urgency of improving surveillance, diagnostics, and treatment access.
At the same time, advances like 4′-FIU highlight the role of genomics, antiviral research, and global collaboration in addressing endemic diseases.
While the drug is still in the experimental stage and requires human clinical trials, it represents a potential shift in how Lassa fever is treated especially in regions where early intervention is difficult.
Lassa fever remains a persistent and evolving public health challenge in Nigeria. But with improved data tracking and emerging therapeutic innovations, there is a clear pathway toward better outcomes and reduced mortality if investments in research, healthcare systems, and community awareness continue.
Read more on the Lassa fever trend in Nigeria .
A recent study in Nature Scientific Reports provides new insights into the genetic diversity of African populations and its implications for disease research and precision medicine. The research highlights how African genomes carry unique and medically relevant genetic variants, many of which are still underrepresented in global datasets.
One of the most important findings is the high variability of disease-associated genetic markers across African populations. For example, variants linked to protection against severe malaria, such as those in the G6PD gene, show significantly different frequencies across regions. Similarly, the well-known sickle cell mutation (HbS) appears at high frequencies in malaria-endemic regions of West and East Africa but is far less common in Southern Africa.
This variation is not random, it reflects centuries of evolutionary adaptation to environmental pressures such as infectious diseases. Another key example is the APOL1 gene variants, which are associated with increased risk of kidney disease. These variants are more common in parts of West Africa, illustrating how genetic adaptations that once provided survival advantages can also influence modern disease risk.
Why This Matters for Research and Medicine
These findings reinforce a critical gap in global health research: African populations remain underrepresented in genomic studies, despite having the highest genetic diversity worldwide. This lack of representation limits the ability to:
- Accurately identify disease risk factors
- Develop effective diagnostics
- Design targeted therapies
- Advance precision medicine
Without diverse biospecimens and genomic data, researchers risk building solutions that do not fully apply to African populations.
The Role of Biobanking and African-Led Research
To bridge this gap, the study underscores the importance of:
- Expanding genomic databases with African representation
- Strengthening biobanking systems for diverse biospecimens
- Supporting collaborative, African-led research initiatives
Biobanks play a central role by enabling the ethical collection, storage, and sharing of biological samples that reflect real population diversity.
At MyAfroDNA, we are committed to advancing inclusive genomic research by providing:
- Access to diverse African biospecimens
- Ethical biobanking and sample management
- Partnerships with researchers, institutions, and healthcare organizations
We invite researchers, universities, and public health institutions to collaborate with us in building datasets and research that truly reflect African populations.
Inclusive genomics is essential for equitable healthcare. Africa must be represented not approximated in global science.
Read more on this research at nature scientific.

Trusted DNA Testing Services in Nigeria
MyAfroDNA provides reliable and confidential DNA testing services in Nigeria for individuals, families, legal professionals, and institutions. Whether you need a paternity DNA test, immigration DNA test, or other relationship testing services, our process is handled with professionalism and strict laboratory standards.
We serve clients across Lagos, Abuja, Port Harcourt, Rivers State, and other cities nationwide.
If you are searching for where to get a DNA test done in Nigeria, MyAfroDNA offers trusted and accurate solutions.
Paternity DNA Testing in Nigeria
Our paternity DNA test determines biological fatherhood with over 99.99% accuracy. This service is available for:
- Peace-of-mind testing
- Legal and court-admissible testing
- Family confirmation purposes
Each test follows secure sample collection procedures and professional laboratory processing to ensure reliable and confidential results.
MyAfroDNA provides paternity DNA testing services across Nigeria with discretion and scientific precision.
Immigration & Legal DNA Testing
MyAfroDNA provides immigration DNA testing services for embassy and visa requirements. Our documentation process follows internationally recognized standards to ensure accurate and legally acceptable results.
We assist families and individuals throughout Nigeria who require DNA testing for official purposes.
Sibling & Relationship DNA Testing
In addition to paternity testing, we provide:
- Sibling DNA testing
- Maternity testing
- Grandparent DNA testing
- Other biological relationship verification services
All tests are conducted using scientifically validated procedures to ensure high levels of accuracy.
Why Choose MyAfroDNA for DNA Testing in Nigeria?
- Accurate and reliable DNA test results
- Strict confidentiality and data protection
- Nationwide service coverage
- Professional laboratory standards
- Efficient turnaround times
- Experienced scientific team
MyAfroDNA combines biotechnology expertise with accessible DNA testing services tailored to individuals and institutions across Nigeria.
DNA Testing Available Nationwide
We provide DNA testing services in:
- Port Harcourt
- Lagos
- Abuja
- Rivers State
- Other locations across Nigeria
If you are searching for “DNA test near me” in Nigeria, our team is available to guide you through the process professionally and confidentially.
Frequently Asked Questions
How much does a DNA test cost in Nigeria?
Our DNA Paternity test is the cheapest in Nigeria and costs 150,000 naira only. Additional charges may apply for shipment of samples or the type of test required. Contact MyAfroDNA directly for current pricing and consultation.
How long does a paternity DNA test take in Nigeria?
Our turnaround time is seven days and can be expedited in cases of emergencies. Our team provides clear timelines during bookings.
Is DNA testing confidential?
Yes. All DNA testing services at MyAfroDNA are handled with strict confidentiality and data protection standards.
Book a DNA Test in Nigeria
To schedule your DNA test appointment, contact MyAfroDNA today.
Call us on 07010644048
Or send us a WhatsApp message on 07010644048 to book an appointment
Our team will guide you through the process professionally and d
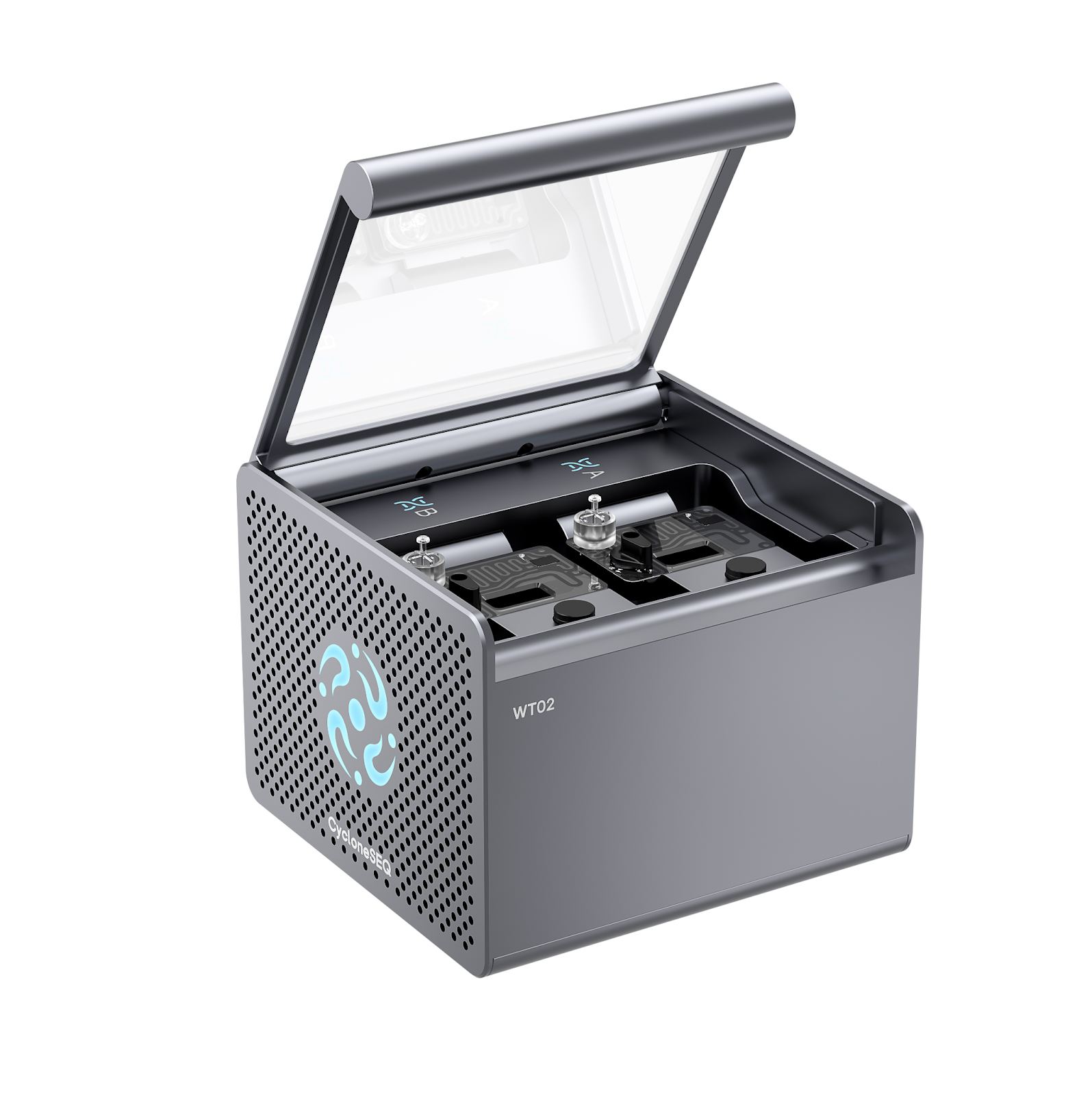

Recently, MyAfroDNA announced the acquisition of the CycloneSEQ sequencer nanopore genome sequencing platform, becoming the first facility in West Africa to deploy this advanced technology. This milestone represents a significant advancement for genome sequencing in Nigeria, biotechnology in West Africa, and African-led genomics research.
But what exactly is the CycloneSEQ platform? How does nanopore sequencing work? And why does this matter for researchers, institutions, and biotech organizations across Africa and globally?
In this article, we explain:
- What the CycloneSEQ platform does
- The science behind its enhanced protein engineering and flow cell design
- Its advantages over traditional sequencing systems
- Why long-read sequencing is critical for modern genomics
- How institutions can collaborate with MyAfroDNA
What Is the CycloneSEQ Nanopore Sequencing Platform?
The CycloneSEQ platform, including the CycloneSEQ-WT02 and CycloneSEQ-WY01 models, is an advanced nanopore-based genome sequencing system developed by MGI Tech Co., Ltd. Unlike conventional short-read sequencing platforms that fragment DNA into small pieces for computational assembly, nanopore sequencing reads long DNA molecules directly as they pass through nanoscale protein pores.
As DNA passes through each nanopore, changes in electrical current are detected at picoampere resolution. These signals are then decoded in real time using advanced basecalling algorithms.
This enables:
- Long-read genome sequencing
- Real-time data generation
- Structural variant detection
- Whole genome assembly
- Metagenomics and pathogen surveillance
- Agricultural and biodiversity genomics
For biotechnology in Nigeria and West Africa, this introduces a new level of sequencing capability previously unavailable locally.
Advanced Technology Behind CycloneSEQ
The CycloneSEQ platform integrates several major innovations that elevate its performance in genomic applications.
Enhanced Protein Engineering
CycloneSEQ utilizes:
- A high thermal stability motor protein
- A structurally rigid nanopore protein
This refined structure improves:
- DNA unwinding efficiency
- Nucleic acid capture
- Reduced sequencing noise
- Signal stability
These enhancements increase threading precision as DNA passes through the pore, significantly improving sequencing reliability.
High-Density Flow Cell Architecture
Each flow cell supports up to 4,096 nanopore protein sensors arranged in a high-density array.
The integration of:
- BioMEMS microfluidic control systems
- ASIC-based electronic signal processing
Allows ultra-sensitive detection of current changes at picoampere levels.
This enables:
- Stable, ultra-long sequencing runs
- High-throughput genomic analysis
- Multi-flow cell scalability
- Continuous real-time sequencing
Optimized Basecalling Algorithm (97% Accuracy)
CycloneSEQ employs a proprietary basecalling algorithm trained using large-scale distributed computing.
With reported accuracy rates of approximately 97%, the system delivers:
- Real-time DNA sequence decoding
- Reliable variant detection
- Reduced computational post-processing
- Improved structural variant resolution
This level of optimization is critical for clinical research, agricultural genomics, and biodiversity projects.
Innovative Nanopore Local Chemistry
A novel sequencing chemistry modulates magnesium ion electrophoretic migration to regulate motor protein activity.
This chemistry improves:
- DNA translocation control
- Sequencing efficiency
- Signal clarity
- Run stability
Together, these innovations position CycloneSEQ as a highly competitive long-read sequencing platform.
CycloneSEQ vs Traditional Sequencing Platforms
Understanding the distinction between long-read nanopore sequencing and traditional short-read systems is essential.
1. Read Length
Traditional platforms generate short DNA fragments requiring computational reassembly.
CycloneSEQ reads long continuous DNA strands, enabling more accurate genome assembly.
2. Structural Variant Detection
Short-read sequencing may struggle with:
- Large insertions
- Repeat expansions
- Complex rearrangements
Long-read sequencing excels in resolving these genomic features.
3. Infrastructure and Accessibility
Traditional systems often require:
- Large-scale laboratory infrastructure
- High capital investment
- Significant operational overhead
CycloneSEQ offers greater flexibility and accessibility for emerging genomics laboratories.
4. Turnaround Time
Outsourcing sequencing abroad can delay research timelines.
With local genome sequencing now available in Nigeria at MyAfroDNA, turnaround time is significantly reduced.
5. Data Sovereignty
Local sequencing strengthens data ownership and supports African-led genomics research initiatives.
Why This Matters for Biotechnology in Nigeria and Africa
Africa contains the highest human genetic diversity globally, yet remains underrepresented in genomic databases.
The deployment of West Africa’s first CycloneSEQ nanopore sequencer strengthens:
- Precision medicine research
- Agricultural biotechnology
- Crop genome assembly
- Pathogen genomics
- Biodiversity and conservation research
- Academic and institutional genomics programs
By expanding DNA sequencing services in Nigeria, MyAfroDNA contributes to positioning Africa as an active contributor, not just a participant, in global genomics.
Frequently Asked Questions
What is nanopore sequencing?
Nanopore sequencing reads DNA directly as it passes through a protein pore, detecting electrical current changes to determine sequence information.
Is CycloneSEQ suitable for whole-genome sequencing?
Yes. It supports long-read whole-genome assembly and structural variant analysis.
Who can partner with MyAfroDNA?
Universities, biotech startups, hospitals, agricultural institutes, NGOs, pharmaceutical researchers, and international collaborators.
Is this technology only for human genomics?
No. It supports plant, microbial, animal, environmental, and metagenomic sequencing applications.
How can organizations access this service?
Institutions can contact MyAfroDNA to discuss project scope, sample type, volume requirements, and turnaround expectations.
Partner with MyAfroDNA
Partner with West Africa’s first nanopore sequencing facility.
MyAfroDNA now offers advanced genome sequencing services in Nigeria, serving researchers and institutions across West Africa, Africa, and globally.
If your organization requires:
- Long-read genome sequencing
- Structural variant analysis
- Agricultural genomics
- Pathogen surveillance
- Biodiversity sequencing
- Precision medicine research support
Our team is ready to collaborate.
Contact MyAfroDNA today to discuss your sequencing project and explore how CycloneSEQ technology can support your research objectives.

Cancer research is entering a transformative era. Precision oncology, the approach of tailoring cancer treatments based on a patient’s unique genetic profile, is already revolutionizing care in many parts of the world. However, recent evidence underscores a critical gap: precision oncology in Africa cannot succeed without African‑centric genomic data, infrastructure, and research partnerships. (Nature Africa, 2026)
Many of the genomic datasets driving current precision oncology tools are largely derived from populations outside Africa. While these datasets have enabled breakthroughs in diagnostics, therapy selection, and prognostic assessments, they do not capture the full spectrum of genetic diversity found in African populations. This matters because cancer development, mutation patterns, and drug responses are influenced by ancestry-specific genomic variations. Without representative African data, treatments risk being less effective or missing critical mutation patterns prevalent in African patients.
Why African Genomes Are Essential for Precision Oncology
The Nature Africa feature emphasizes that African populations possess high genetic diversity, one of the richest in the world. This diversity affects not only susceptibility to certain cancer types but also the effectiveness of targeted therapies. For example, mutations in genes such as TP53, BRCA1/2, or EGFR may appear with different frequencies or impact drug responses differently in African patients compared to non-African populations. Precision oncology that ignores this variation risks exacerbating health disparities, rather than reducing them.
The Role of Biobanking and Genomic Research
To build effective precision oncology for Africa, robust biobanks and genomic infrastructure are vital. Biobanks store ethically collected biological samples — including blood, tissue, saliva, and DNA — under controlled conditions for research. They allow researchers to:
- Identify population-specific genetic variants
- Map tumor evolution in African patients
- Develop diagnostic assays tailored to local mutation patterns
- Support clinical trials that reflect real-world African populations
Ethical stewardship and consent are central to biobanking. Samples must be traceable, stored securely, and used transparently for approved research. African biobanks like MyAfroDNA aim to provide this foundation while supporting collaborations with hospitals, universities, and research institutions.
Investing in African Genomic Capacity
Precision oncology requires more than samples; it needs trained personnel, advanced sequencing technologies, molecular tumor boards, and data analysis capacity. African researchers and clinicians must be empowered to interpret genomic information in context, linking mutations to effective therapies. Collaborative networks allow the sharing of resources, expertise, and knowledge to accelerate the translation of genomic insights into patient care.
Partner With MyAfroDNA for Precision Oncology for Africans by Africans
At MyAfroDNA, we are committed to advancing African-centric precision oncology through:
- Ethical biospecimen collection and storage
- Access to diverse African genomic samples for research
- Partnerships with researchers, hospitals, and institutions
- Training programs in genomic analysis and precision medicine
We invite researchers, universities, hospitals, and pharmaceutical companies to collaborate with us. By working together, we can ensure that precision oncology in Africa is informed by representative genomic data, supports tailored therapies, and contributes to equitable healthcare outcomes.
African genomic research is not just a scientific opportunity; it is a responsibility. Every sample, study, and collaboration strengthens the foundation for cancer care that is designed by Africans, for Africans.
Partner with MyAfroDNA today to advance precision oncology and genomic research in Africa.
Source: Nature Africa — Precision oncology specifically for Africa (nature.com)
MyAfroDNA, an African biotechnology company focused on advancing genomics research and innovation on the African continent, has announced the acquisition and deployment of the MGI CycloneSEQ™ genome sequencer, becoming the first facility in West Africa to deploy this advanced, portable sequencing platform.
CycloneSEQ is a cost-efficient long-read sequencing platform suitable for generating high-quality reference genomes across diverse biological systems. Its accuracy, scalable throughput, and streamlined workflows support reliable genome sequencing, assembly, and variant resolution, enabling large-scale sequencing projects with reduced per-sample cost compared to conventional long-read approaches. The system’s compact footprint and stable operation make it appropriate for sustained deployment in African research environments, facilitating locally led genome projects while strengthening regional capacity and data stewardship.
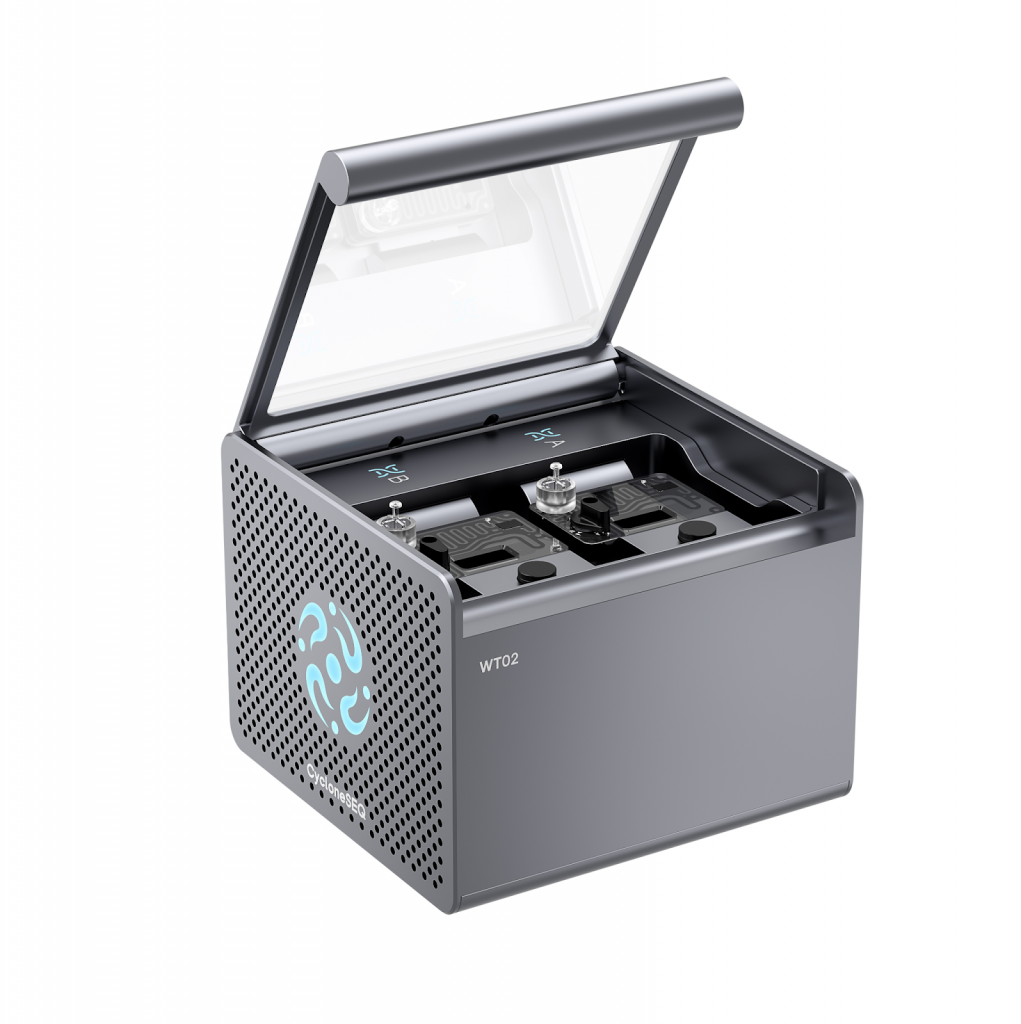

Figure 1: Photo of recently acquired CycloneSEQ genome sequencer deployed at MyAfroDNA facility in Port Harcourt, Nigeria
The CycloneSEQ™ significantly strengthens MyAfroDNA’s capacity to perform high-resolution, high-accuracy genomic sequencing, enabling applications across human health, agriculture, biodiversity, and environmental research. This milestone marks a major step toward expanding Africa’s ability to generate and analyze genomic data locally, reducing reliance on external sequencing infrastructure.
“The acquisition of the CycloneSEQ™ genome sequencer significantly strengthens our ability to generate high-quality whole-genome data locally and at scale. It allows us to support research across health, agriculture, and biodiversity, while ensuring African-led projects are powered by African infrastructure, expertise, and data ownership,” said Dr. Justin Eze Ideozu, Founder of MyAfroDNA.
Designed for precision and scalability, the CycloneSEQ™ supports whole-genome sequencing, reference genome development, population genomics, and accurate detection of genetic variants. Its flexible workflows enable MyAfroDNA to support both targeted studies and large-scale genomic projects requiring high data quality.
As part of this deployment, MyAfroDNA will immediately begin generating high-quality reference genomes to support biodiversity and conservation genomics in partnership with the African BioGenome Project (AfricaBP) and the West Africa Regional Node of the AfricaBP, Regional Center for Biotechnology and Bioresources Research (RCBBR), University of Port Harcourt, Nigeria.
“The presence of CycloneSEQ™ at MyAfroDNA is a significant development which will certainly inject life into the Endangered and Endemic Species BioGenome Project of the West African Regional node of AfricaBP. It will, as well, facilitate realization of the AfricaBP-10KP 2.0 and AfricaBP Plant Genome Projects. Hopefully, our first genome sequence will be rolled out soon. I sincerely congratulate Dr Justin Eze Ideozu and MyAfroDNA for achieving this great milestone”, said Professor Julian Osuji, AfricaBP West Africa Regional Node Coordinator and Director of the Regional Center for Biotechnology and Bioresources Research (RCBBR), University of Port Harcourt, Nigeria.
“We are happy to see the CycloneSEQ™ deployed in West Africa with MyAfroDNA. This installation represents what MGI aims to achieve globally - democratizing access to advanced sequencing technologies. We look forward to supporting MyAfroDNA and AfricaBP in their mission to sequence Africa’s endemic species by placing high-accuracy long-read capabilities directly into the hands of African researchers", said Chen Fang, General Manager of MGI Europe and Africa.
“MyAfroDNA and MGI are two strong partners of the African BioGenome Project (AfricaBP) who have demonstrated strong commitments to the African genomics and molecular biology landscape. To sequence African indigenous and endemic biological species will require intentional investments by African organisations, and I’m quite pleased (and particularly proud) to see MyAfroDNA take this bold step in acquiring the portable CycloneSEQ genome sequencer. This is a testament to the AfricaBP ecosystem in enabling partnerships, collaborations, and local investments, adding to its progress of consistently provoking local actions. The AfricaBP cannot wait to kickstart its sequencing with MyAfroDNA, and supporting biodiversity genome sequencing in Africa”, said Dr. ThankGod Echezona Ebenezer, Founder and Co-Chair, AfricaBP.
This acquisition positions West Africa as an active contributor to global genomics and life sciences research. By enabling in-region sequencing, MyAfroDNA will support universities, research institutes, conservation organizations, biotechnology companies, and policymakers with access to advanced genomic infrastructure previously unavailable in the region.
This development reinforces MyAfroDNA’s broader mission to strengthen Africa’s biotechnology ecosystem and apply genomics to real-world challenges in health, agriculture, biodiversity conservation, and environmental sustainability.
About MyAfroDNA
We are an African biotechnology company that provides solutions to the significant lack of African representation in clinical, genomics, and translational research. Contact us for questions: info@myafrodna.com

A new study published in Nature projects that climate change, especially extreme weather events could lead to a substantial increase in malaria cases and deaths across Africa over the next 25 years.
Using 25 years of climate, health, socioeconomic, and disease control data, researchers estimate that climate change could contribute to up to 123 million additional malaria cases and more than 500,000 additional deaths between 2024 and 2050 if current control strategies remain unchanged.
While many studies have focused on how temperature and rainfall affect mosquito ecology, this research shows that extreme weather such as floods and cyclones — will be the main driver of increased risk, disrupting housing, health services, prevention programs, and access to treatment. These disruptions may account for the majority of additional cases and deaths in the coming decades.
Most of the projected increases will occur in regions where malaria is already endemic rather than in entirely new areas, underscoring the vulnerability of existing control systems to climate shocks. These findings highlight a profound challenge: climate change doesn’t just reshape environments, it can undermine disease control infrastructures, particularly where health systems are already fragile.
For genomic scientists and public health researchers, this study reinforces the urgency of climate-resilient malaria strategies that integrate environmental forecasting, vector surveillance, health system preparedness, and genomic research to track parasite and vector evolution under changing conditions.
At MyAfroDNA, we are committed to supporting collaborative research that advances:Climate-informed malaria surveillance Integrated vector genomics and environmental data systemsCommunity-embedded biospecimen collection and analysisData-driven intervention and control strategiesIf you are a researcher, institution, or organization working on malaria, climate health, genomics, or vector-borne disease research, we invite you to partner with MyAfroDNA. Together, we can develop solutions that are tailored to Africa’s climate and health needs.
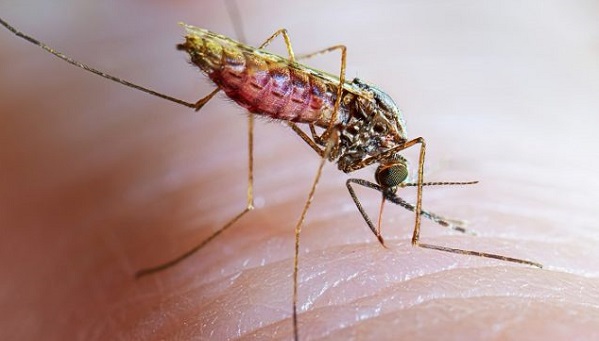
Malaria remains one of Africa’s most persistent public health challenges, and early detection of disease vectors is critical to reducing transmission. A recent development in Madagascar highlights how artificial intelligence (AI) and citizen science are reshaping how malaria threats are identified and monitored across the continent.
According to a report by SciDev.Net, researchers used AI-powered image recognition tools alongside community participation to detect a potentially invasive malaria mosquito species, Anopheles stephensi, in Madagascar. This species is particularly concerning because it thrives in urban environments, breeding in water containers such as buckets, tyres, and tanks, settings where malaria transmission has traditionally been lower.
The detection was triggered when a citizen submitted a mosquito larva image via the NASA GLOBE Observer app. AI models trained on thousands of verified mosquito images classified the specimens with high confidence, demonstrating how digital tools can complement traditional entomological surveillance, especially in resource-limited settings.
For Africa, this approach has major implications. Conventional mosquito surveillance relies heavily on field trapping, microscopy, and laboratory confirmation, methods that are often expensive, slow, and unevenly distributed. AI-supported systems allow for rapid, scalable, and community-driven monitoring, improving early warning capacity and public health response.
At MyAfroDNA, we view this as part of a broader shift toward integrated genomic, digital, and environmental surveillance. Combining AI, biospecimen analysis, and genomic research can strengthen malaria control strategies, support vector mapping, and improve our understanding of how mosquito populations evolve and spread.
MyAfroDNA is actively seeking collaboration with researchers, public health institutions, universities, and malaria-focused organisations to support:
- Vector genomics and biospecimen research
- AI-assisted disease surveillance
- Data-driven malaria prevention strategies
Together, we can build African-led, evidence-based solutions for malaria research and control.
Reach out to collaborate with MyAfroDNA.
Source: SciDev.Net – AI and citizens detect invasive mosquito in Madagascar
https://www.scidev.net/sub-saharan-africa/news/ai-and-citizens-detect-invasive-mosquito-in-madagascar/

Public health surveillance has traditionally relied on clinics, hospitals, and laboratories to report cases of disease. But across much of Africa, this system captures only part of the picture. Many people do not seek care, cannot access testing, or are excluded from formal health systems altogether. As a result, outbreaks are often detected late, under-reported, or missed entirely.
A recent report in Nature highlights how wastewater surveillance is changing this reality, using sewage to track disease trends at the community level, even when clinical data is limited.
What is wastewater surveillance?
Wastewater surveillance involves testing sewage for genetic material from pathogens such as viruses and bacteria. When people are infected, traces of these pathogens are shed through bodily waste and end up in wastewater systems. By analysing these samples, scientists can estimate disease circulation across entire communities, without relying on individual testing or hospital visits.
In practical terms, one wastewater sample can represent thousands or even millions of people.
Lessons from South Africa
Researchers in South Africa demonstrated how wastewater monitoring can reveal disease trends that conventional surveillance fails to capture. By comparing viral signals in sewage with reported clinical cases, they found a significant gap between actual infections and officially recorded data.
In several instances, wastewater data showed rising infection levels weeks before clinics recorded an increase in cases. This early signal is especially important in lower-income or underserved areas, where testing rates tend to be lower, and outbreaks are more likely to go unnoticed.
The findings underscore a critical reality: absence of data does not mean absence of disease.
Beyond COVID-19
While wastewater surveillance gained prominence during the COVID-19 pandemic, its applications extend far beyond one virus. Researchers are now detecting genetic material linked to:
- Measles
- Mpox
- Influenza
- Hepatitis A and E
In some cases, these pathogens were identified in wastewater even when no clinical cases had been officially reported in the area. This positions wastewater surveillance as a powerful early-warning system for emerging and re-emerging diseases.
Why this matters for Africa
Wastewater surveillance offers several advantages that are particularly relevant in African contexts:
- Equity: It captures health data from entire communities, including people who are not represented in clinical systems.
- Cost-effectiveness: Monitoring a few sites can be more affordable than mass individual testing.
- Early detection: Public health authorities can respond sooner, potentially preventing wider outbreaks.
- Genomic insight: Sequencing wastewater samples allows scientists to track variants and pathogen evolution over time.
For regions facing resource constraints, these benefits can strengthen public health decision-making without placing additional burdens on individuals.
The challenges ahead
Despite its promise, wastewater surveillance is not without limitations. It requires:
- Laboratory infrastructure and sequencing capacity
- Skilled personnel for data analysis
- Sustainable funding and policy support
Encouragingly, continental initiatives, including efforts led by Africa CDC, are working toward integrating wastewater and environmental surveillance into broader disease monitoring systems.
Where MyAfroDNA fits in
At MyAfroDNA, we believe that genomic tools should serve real-world public health needs, especially in underrepresented regions. Wastewater surveillance demonstrates how genomics can move beyond laboratories and clinics to inform population-level health decisions.
As Africa continues to invest in biospecimen science, molecular testing, and genomic research, approaches like wastewater surveillance will be essential for building resilient, inclusive health systems.
Better data leads to better decisions and better outcomes for communities.

