Recently, MyAfroDNA announced the acquisition of the CycloneSEQ sequencer nanopore genome sequencing platform, becoming the first facility in West Africa to deploy this advanced technology. This milestone represents a significant advancement for genome sequencing in Nigeria, biotechnology in West Africa, and African-led genomics research.
But what exactly is the CycloneSEQ platform? How does nanopore sequencing work? And why does this matter for researchers, institutions, and biotech organizations across Africa and globally?
In this article, we explain:
- What the CycloneSEQ platform does
- The science behind its enhanced protein engineering and flow cell design
- Its advantages over traditional sequencing systems
- Why long-read sequencing is critical for modern genomics
- How institutions can collaborate with MyAfroDNA
What Is the CycloneSEQ Nanopore Sequencing Platform?
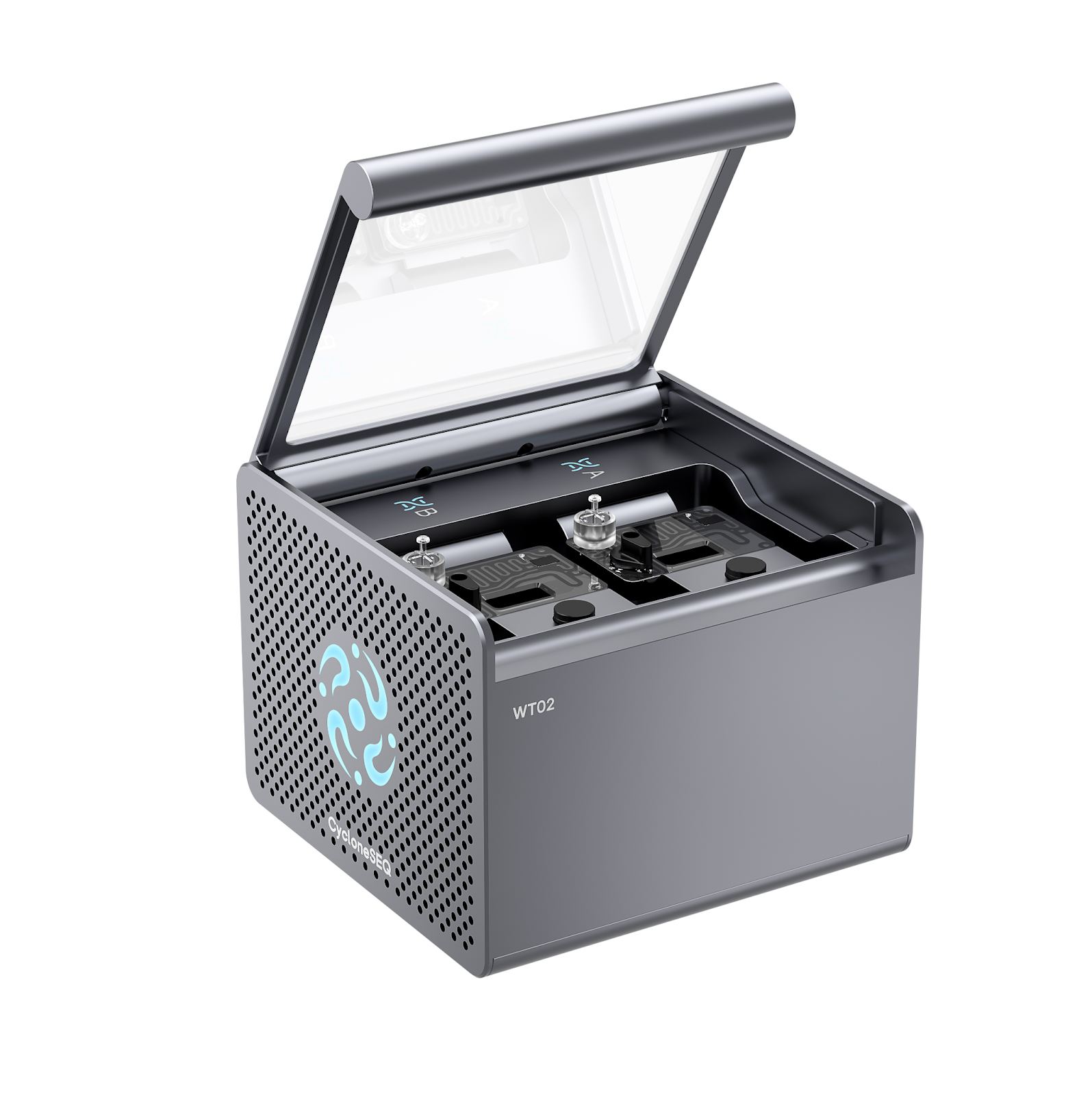
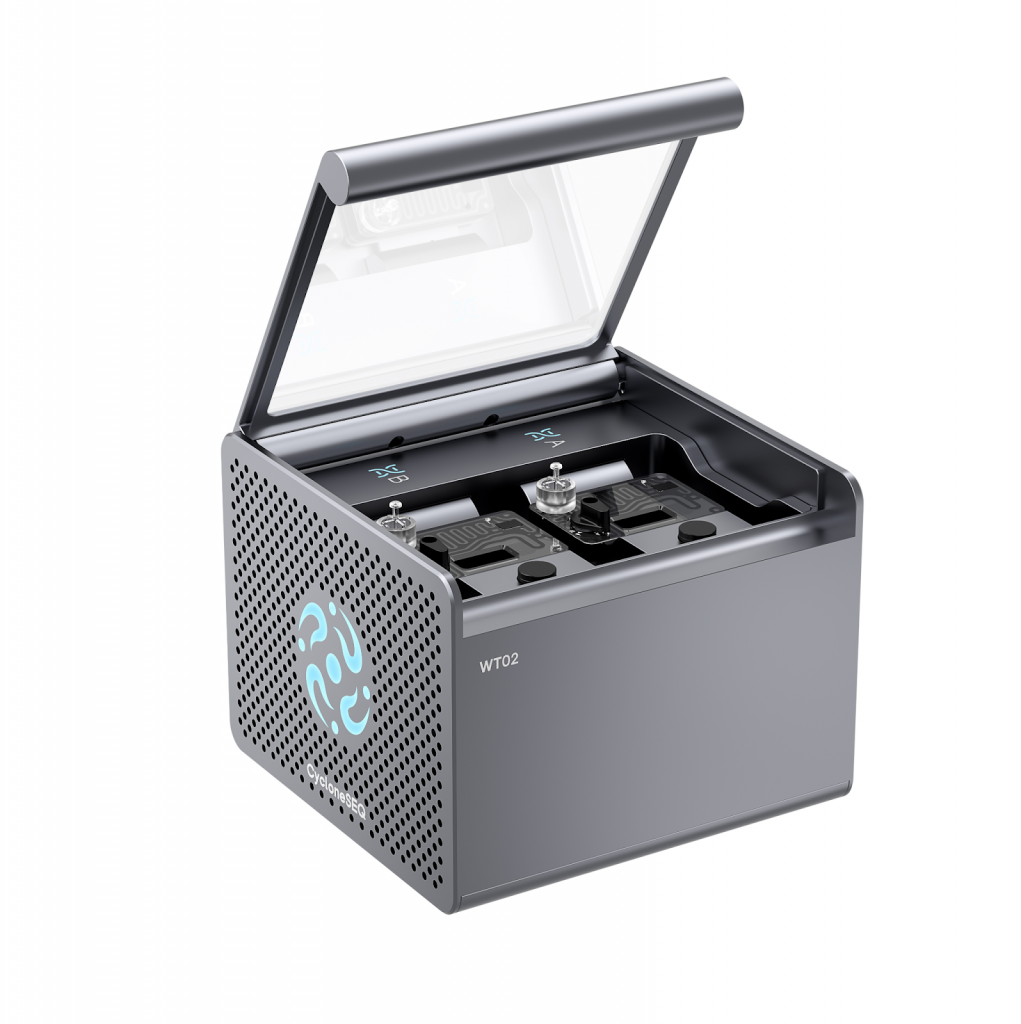
The CycloneSEQ platform, including the CycloneSEQ-WT02 and CycloneSEQ-WY01 models, is an advanced nanopore-based genome sequencing system developed by MGI Tech Co., Ltd. Unlike conventional short-read sequencing platforms that fragment DNA into small pieces for computational assembly, nanopore sequencing reads long DNA molecules directly as they pass through nanoscale protein pores.
As DNA passes through each nanopore, changes in electrical current are detected at picoampere resolution. These signals are then decoded in real time using advanced basecalling algorithms.
This enables:
- Long-read genome sequencing
- Real-time data generation
- Structural variant detection
- Whole genome assembly
- Metagenomics and pathogen surveillance
- Agricultural and biodiversity genomics
For biotechnology in Nigeria and West Africa, this introduces a new level of sequencing capability previously unavailable locally.
Advanced Technology Behind CycloneSEQ
The CycloneSEQ platform integrates several major innovations that elevate its performance in genomic applications.
Enhanced Protein Engineering
CycloneSEQ utilizes:
- A high thermal stability motor protein
- A structurally rigid nanopore protein
This refined structure improves:
- DNA unwinding efficiency
- Nucleic acid capture
- Reduced sequencing noise
- Signal stability
These enhancements increase threading precision as DNA passes through the pore, significantly improving sequencing reliability.
High-Density Flow Cell Architecture
Each flow cell supports up to 4,096 nanopore protein sensors arranged in a high-density array.
The integration of:
- BioMEMS microfluidic control systems
- ASIC-based electronic signal processing
Allows ultra-sensitive detection of current changes at picoampere levels.
This enables:
- Stable, ultra-long sequencing runs
- High-throughput genomic analysis
- Multi-flow cell scalability
- Continuous real-time sequencing
Optimized Basecalling Algorithm (97% Accuracy)
CycloneSEQ employs a proprietary basecalling algorithm trained using large-scale distributed computing.
With reported accuracy rates of approximately 97%, the system delivers:
- Real-time DNA sequence decoding
- Reliable variant detection
- Reduced computational post-processing
- Improved structural variant resolution
This level of optimization is critical for clinical research, agricultural genomics, and biodiversity projects.
Innovative Nanopore Local Chemistry
A novel sequencing chemistry modulates magnesium ion electrophoretic migration to regulate motor protein activity.
This chemistry improves:
- DNA translocation control
- Sequencing efficiency
- Signal clarity
- Run stability
Together, these innovations position CycloneSEQ as a highly competitive long-read sequencing platform.
CycloneSEQ vs Traditional Sequencing Platforms
Understanding the distinction between long-read nanopore sequencing and traditional short-read systems is essential.
1. Read Length
Traditional platforms generate short DNA fragments requiring computational reassembly.
CycloneSEQ reads long continuous DNA strands, enabling more accurate genome assembly.
2. Structural Variant Detection
Short-read sequencing may struggle with:
- Large insertions
- Repeat expansions
- Complex rearrangements
Long-read sequencing excels in resolving these genomic features.
3. Infrastructure and Accessibility
Traditional systems often require:
- Large-scale laboratory infrastructure
- High capital investment
- Significant operational overhead
CycloneSEQ offers greater flexibility and accessibility for emerging genomics laboratories.
4. Turnaround Time
Outsourcing sequencing abroad can delay research timelines.
With local genome sequencing now available in Nigeria at MyAfroDNA, turnaround time is significantly reduced.
5. Data Sovereignty
Local sequencing strengthens data ownership and supports African-led genomics research initiatives.
Why This Matters for Biotechnology in Nigeria and Africa
Africa contains the highest human genetic diversity globally, yet remains underrepresented in genomic databases.
The deployment of West Africa’s first CycloneSEQ nanopore sequencer strengthens:
- Precision medicine research
- Agricultural biotechnology
- Crop genome assembly
- Pathogen genomics
- Biodiversity and conservation research
- Academic and institutional genomics programs
By expanding DNA sequencing services in Nigeria, MyAfroDNA contributes to positioning Africa as an active contributor, not just a participant, in global genomics.
Frequently Asked Questions
What is nanopore sequencing?
Nanopore sequencing reads DNA directly as it passes through a protein pore, detecting electrical current changes to determine sequence information.
Is CycloneSEQ suitable for whole-genome sequencing?
Yes. It supports long-read whole-genome assembly and structural variant analysis.
Who can partner with MyAfroDNA?
Universities, biotech startups, hospitals, agricultural institutes, NGOs, pharmaceutical researchers, and international collaborators.
Is this technology only for human genomics?
No. It supports plant, microbial, animal, environmental, and metagenomic sequencing applications.
How can organizations access this service?
Institutions can contact MyAfroDNA to discuss project scope, sample type, volume requirements, and turnaround expectations.
Partner with MyAfroDNA
Partner with West Africa’s first nanopore sequencing facility.
MyAfroDNA now offers advanced genome sequencing services in Nigeria, serving researchers and institutions across West Africa, Africa, and globally.
If your organization requires:
- Long-read genome sequencing
- Structural variant analysis
- Agricultural genomics
- Pathogen surveillance
- Biodiversity sequencing
- Precision medicine research support
Our team is ready to collaborate.
Contact MyAfroDNA today to discuss your sequencing project and explore how CycloneSEQ technology can support your research objectives.